Table of contents
Structure search Help
Monomeric structure description
In Norine, we use monomeric structures to represent the nonribosomal peptides. These structures are graphs with
nodes labelled by monomers. We adopt a convention to write those graphs as formatted strings.
We use 3 letter codes for the classical amino acids and add the additional groups for their derivatives such as OH-Asp (Hydroxyaspartic acid) for Asp (Aspartic Acid).
We also design short names for the other compounds. The complete monomer list can be consulted here.
You can search for a structural pattern in all peptides of Norine. The pattern must be specified by a monomeric structure. In addition to the usual graph representations, a structural pattern may have nodes labelled by several alternative monomers. These can be specified by
- a list a monomers between brackets, such as [Ala|Ser] that means that either Ala or Ser can be at the corresponding position
- a cluster name, such as Ala* that means all the derivatives of Alanine. You have all the clusters with their monomers in this page. If monomers in your peptide are not in the list, you can use the name of the cluster that should have contained it instead.
- one or several clusters can be added in a list between brackets
- the joker symbol, 'X', stands for any monomer. We also provide the 'D-X' joker for any D-monomer.
| vancomycin | ||||||||
| 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 |
| Asn | bOH-Cl-Tyr | NMe-Leu | Hpg | D-Glc | Van | bOH-Cl-Tyr | Dhpg | Hpg |
| @1,3 | @0,2,3 | @1 | @1,0,4,6,8 | @3,5 | @4 | @3,7,8 | @6,8 | @3,7,6 |
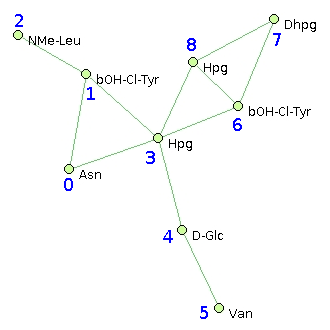
Such graphs can be either obtained from Norine peptide description pages (by copy-paste the "graph representation" lines), or drawn using the dedicated structure editor (consult its help page).
Monomer composition fingerprint search (MCFP)
MCFP is represented as an integer vector, in which each element represents the presence (number of occurrences) or
absence ("0" value) of a specific monomer. The process of generating the MCFP for each peptide starts by extracting the
monomer compositions from Norine and then filling the corresponding positions in the MCFP vector. A second vector is added from
the structure of the peptide, each value is the number of nomoner for a degree value (number of link).
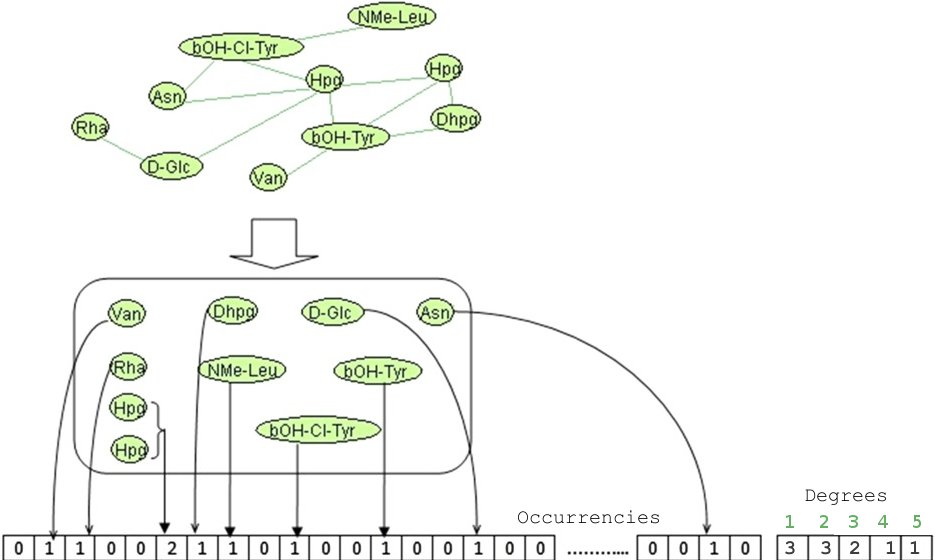
We use Tanimoto-based similarity search system (TAN). This system is based on the Tanimoto coefficient that is a well
established method in similarity-based virtual screening and was therefore used as reference. In particular, the
continuous form of the Tanimoto coefficient was used. If Ai and Bi
represent the ith monomer occurrence and Adegj and Bdegj represents the number of monomer of degree j in the peptides
A and B, respectively,
then the similarity score SA,B between peptides A and B was calculated by the following equation.
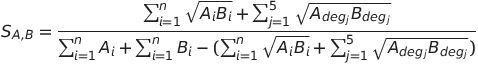
The TAN coefficient varies between 0 (totally different monomer compositions) and 1 (identical monomer compositions). For more information, please consult this article "A new fingerprint to predict nonribosomal peptides activity" (Journal of Computer-Aided Molecular Design, 2012, Vol 26(10), pp 1187-1194, A. Abdo,S. Caboche, V. Leclere, P. Jacques, M. Pupin ).
Structural pattern search
A pattern **occurs** in a peptide graph if it **occurs** as a subgraph, such that each node of the pattern labelled by a monomer is associated to a node of the peptide labelled by the same monomer, and a node labelled by alternative monomers is associated to one of them.
For example, consider the following set of peptide structures :- Ala, Val, His, Pro-Thz, D-Leu @1 @0,2 @1,3 @2,4 @3
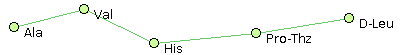
- Ala, Val, Gly, Pro-Thz, D-Leu @1 @0,2 @1,3 @2,4 @3
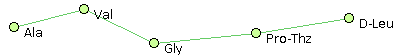
- Glu, Val, His, Pro-Thz, D-Leu @1,4 @0,2 @1,3 @2,4 @0,3
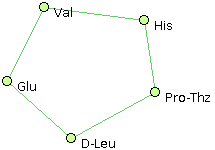
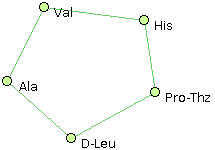
- Ala, Val, His, Pro-Thz, D-Leu @1,2 @0,2 @0,1,3 @2,4 @3

- Val, His, Pro-Thz, D-Leu, Ala @1,4 @0,2 @1,3 @2,4 @0,3
- The structural pattern Ala, Val, His @1 @0,2 @1 matches structures 1, 4 and 5.
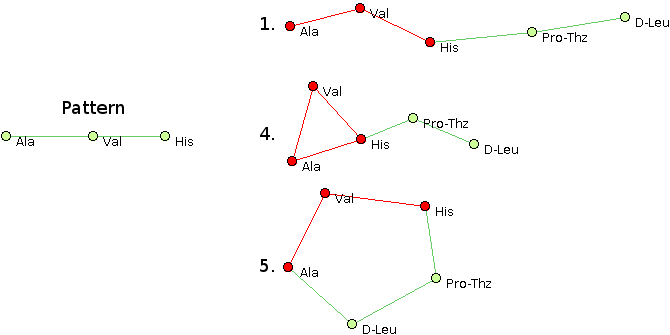
- The structural pattern Ala, Val, X @1 @0,2 @1 matches
structures 1, 2, 4 and 5.
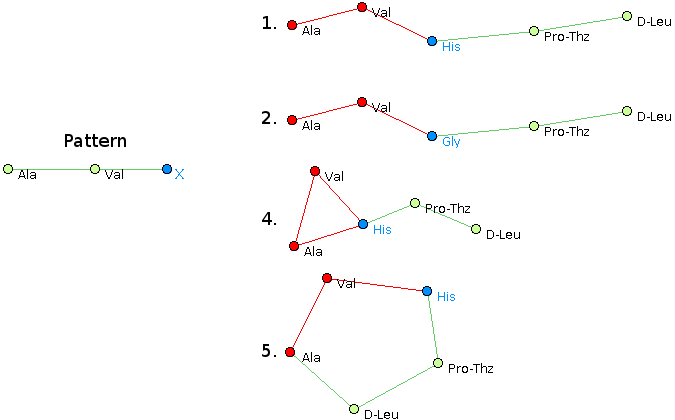
- The latter result can also be obtained with pattern Ala, Val, [His|Gly] @1 @0,2 @1.
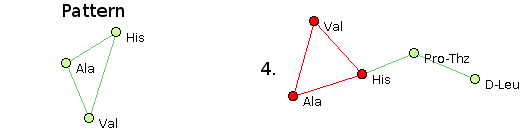
- The structural pattern Ala, Val, His @1,2 @0,2 @0,1 matches structure 4 only.
Similarity search
Similarity search allows the user to search for peptides which have a similar monomeric structure than the query
structure. For each peptide of the Norine database, a distance between the query structure and the query is calculated.
The distance ranges from 0, identical strucutures to 1, totally different structures.
The proposed clustering levels are the one presented in the monomers list. No
clustering means that only monomers are considered, clustering 1 stands for the inner level of clustering, for
example Ala* and clustering 2 for the outer level, for example Ala+Gly.